[1] 0.61 Today’s Lecture Objectives
Bayes’ Theorem
Likelihood function, Posterior distribution
How to report posterior distribution of parameters
Bayesian update
but, before we begin…
2 Refresh the memory: What is Bayesian?
- What are key components of Bayesian models?
- Likelihood function - data distribution
- Prior distribution - belief / previous evidences of parameters
- Posterior distribution - updated information of parameters given our data and model
- Posterior predictive distribution - future / predicted data
- What are the differences between Bayesian with Frequentist analysis?
- prior distribution: Bayesian
- hypothesis of fixed parameters: frequentist
- estimation process: MCMC vs. Maximum likelihhood estimation (MLE) or ordinary least square (OLS)
- posterior distribution vs. point estimates of parameters
- credible interval (plausibility of the parameters having those values) vs. confidence interval (the proportion of infinite samples having the fixed parameters)
3 Bayes’ Theorem: How Bayesian Statistics Work
Bayesian methods rely on Bayes’ Theorem
P(\theta | Data) = \frac{P(Data|\theta)P(\theta)}{P(Data)} \propto P(Data|\theta) P(\theta)
Where:
- P: probability distribution function (PDF)
- P(\theta|Data) : the posterior distribution of parameter \theta, given the observed data
- P(Data|\theta): the likelihood function (conditional distributin) of the observed data, given the parameters
- P(\theta): the prior distribution of parameter \theta
- P(Data): the marginal distribution of the observed data
4 Preparation: Install CmdStan
5 Example One: Six-sided die
Suppose we want to assess the probability of rolling a “1” on a six-sided die:
\theta \equiv p_{1} = P(D = 1)
Suppose we collect a sample of data (N = 5):
Data \equiv X = \{0, 1, 0, 1, 1\}The prior distribution of parameters is denoted as P(p_1) ;
The likelihood functionis denoted as P(X | p_1);
The posterior distribution is denoted as P(p_1|X)
Then we have:
P(p_1|X) = \frac{P(X|p_1)P(p_1)}{P(X)} \propto P(X|p_1) P(p_1)
6 Baysian Inference Diagram
7 Step 1: Choose the Likelihood function (Model)
The Likelihood function P(X|p_1) follows Binomial distribution of 3 succuess out of 5 samples:
P(X|p_1) = \prod_{i =1}^{N=5} p_1^{X_i}(1-p_1)^{X_i} \\= (1-p_i) \cdot p_i \cdot (1-p_i) \cdot p_i \cdot p_iQuestion here: WHY use Bernoulli (Binomial) Distribution (feat. Jacob Bernoulli, 1654-1705)?
My answer: Bernoulli dist. has nice statistical probability. “Nice” means making totally sense in normal life– a common belief. For example, the p_1 value that maximizes the Bernoulli-based likelihood function is Mean(X), and the p_1 values that minimizes the Bernoulli-based likelihood function is 0 or 1
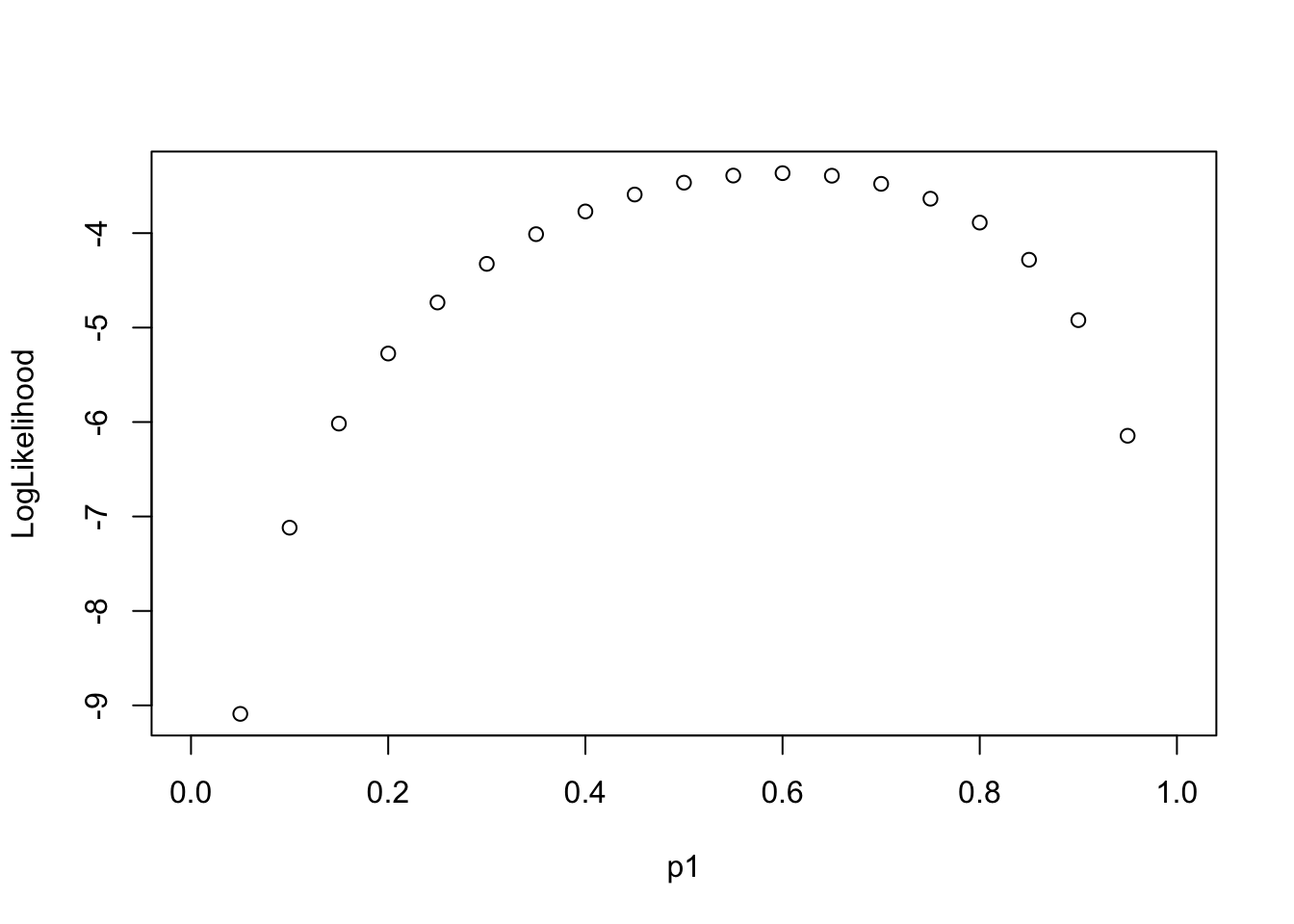
7.1 Log-likelihood Function along “Parameter”
Assume each event is independent, the multiplication of the probabilities of each event is the “Likelihood”. Log transform of Likelihood is easier for further analysis.
LL(p_1|\{0, 1, 0, 1, 1\})
Goal: find the most possible p1 value (the probability of the event) so that the data {0, 1, 0, 1, 1} has the highest likelihood
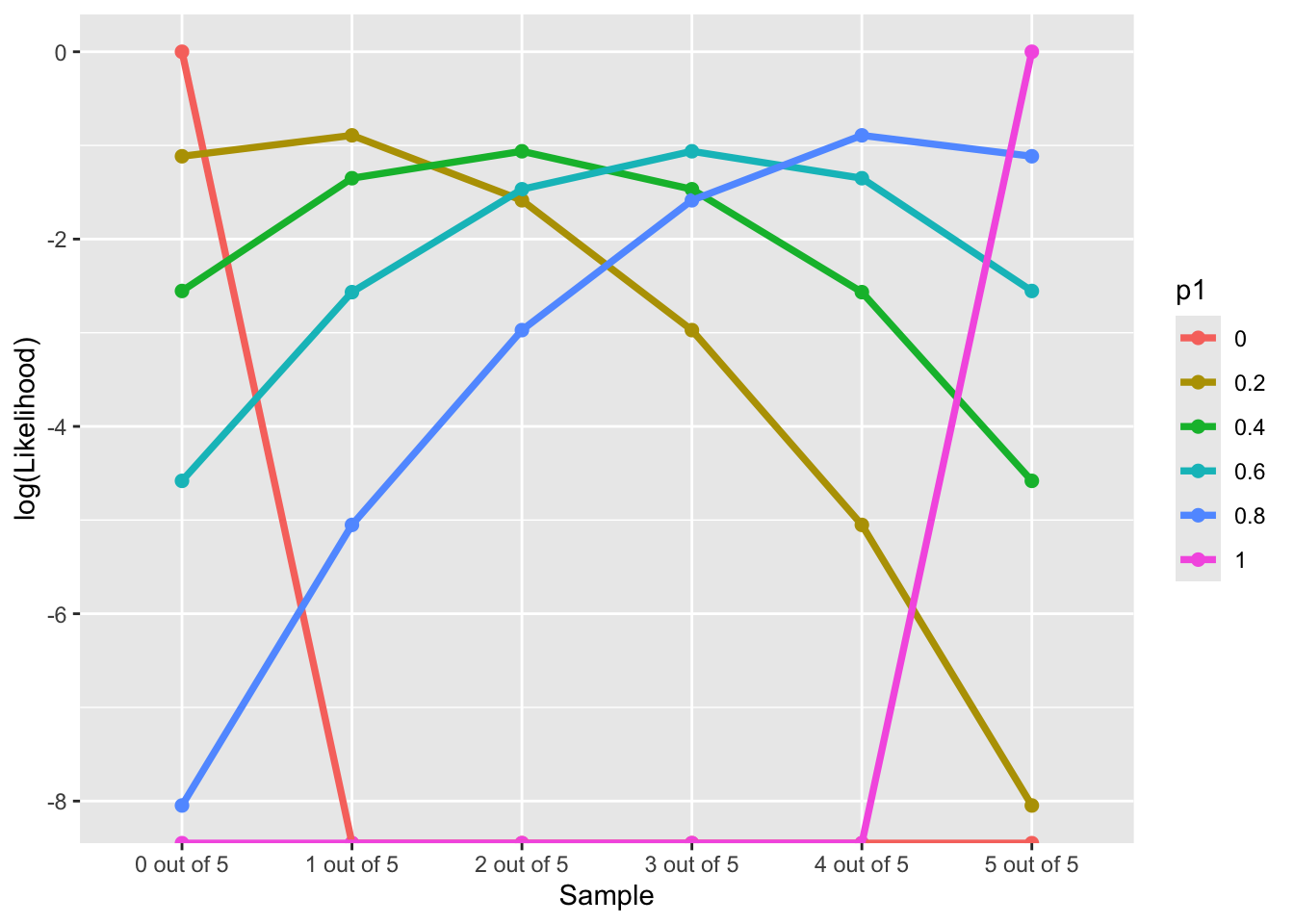
7.2 Log-likelihood Function along “Data”
LL(Data|p_1 \in \{0, 0.2,0.4, 0.6, 0.8,1\})
Click here to see R code
library(tidyverse)
p1 = c(0, 2, 4, 6, 8, 10) / 10
nTrails = 5
nSuccess = 0:nTrails
Likelihood = sapply(p1, \(x) choose(nTrails,nSuccess)*(x)^nSuccess*(1-x)^(nTrails - nSuccess))
Likelihood_forPlot <- as.data.frame(Likelihood)
colnames(Likelihood_forPlot) <- p1
Likelihood_forPlot$Sample = factor(paste0(nSuccess, " out of ", nTrails), levels = paste0(nSuccess, " out of ", nTrails))
# plot
Likelihood_forPlot %>%
pivot_longer(-Sample, names_to = "p1") %>%
mutate(`log(Likelihood)` = log(value)) %>%
ggplot(aes(x = Sample, y = `log(Likelihood)`, color = p1)) +
geom_point(size = 2) +
geom_path(aes(group=p1), size = 1.3)8 Step 2: Choose the Prior Distribution for p_1
We must now pick the prior distribution of p_1:
P(p_1)
Compared to likelihood function, we have much more choices. Many distributions to choose from
To choose prior distribution, think about what we know about a “fair” die.
the probability of rolling a “1” is about \frac{1}{6}
the probabilities much higher/lower than \frac{1}{6} are very unlikely
Let’s consider a Beta distribution as the prior distribution of p_1:
p_1 \sim Beta(\alpha, \beta)
or
P(p_1) \equiv Beta(\alpha, \beta)
8.1 The Beta Distribution
For parameters that range between 0 and 1, the beta distribution makes a good choice for prior distribution:
P(p_1) = \frac{(p_1)^{a-1} (1-p_1)^{\beta-1}}{B(\alpha,\beta)}
Where denominator, B, is a normalization constant to ensure that the total probability is 1:
B(\alpha,\beta) = \frac{\Gamma(\alpha)\Gamma(\beta)}{\Gamma(\alpha+\beta)}
and (fun fact: derived by Daniel Bernoulli (1700-1782)),
\Gamma(\alpha) = \int_0^{\infty}t^{\alpha-1}e^{-t}dt
8.2 Beta Functions in R
Think about 3 sets of 10 dices. They have varied probability of “1”.
Question1: any fair dice among 30 dices?
Question2: which set have relatively fair dices.
Hint: 1 / 6 = 0.16666...
set.seed(4)
alpha = 2
beta = 2
rbeta(n = 10, shape1 = alpha, shape2 = beta) # randomly generate p1 given beta(2, 2) [1] 0.5609713 0.3497046 0.7392076 0.6644891 0.8877412 0.6888135 0.4400386
[8] 0.9230177 0.9079563 0.5044343alpha = 1
beta = 1
rbeta(n = 10, shape1 = alpha, shape2 = beta) # randomly generate p1 given beta(1, 1) [1] 0.35083863 0.51800100 0.48629826 0.43288779 0.12200407 0.51762911
[7] 0.53997409 0.61158197 0.06168091 0.43440547alpha = 200
beta = 996
rbeta(n = 10, shape1 = alpha, shape2 = beta) # randomly generate p1 given beta(200, 996) [1] 0.1851006 0.1746928 0.1637927 0.1842677 0.1790450 0.1558018 0.1840178
[8] 0.1691043 0.1580080 0.14830288.3 More Beta Distribution
The Beta distribution has a mean of \frac{\alpha}{\alpha+\beta} and a mode of \frac{\alpha -1}{\alpha + \beta -2} for \alpha > 1 & \beta > 1; (fun fact: when \alpha\ \&\ \beta< 1, pdf is U-shape, what that mean?)
\alpha and \beta are called hyperparameters of the parameter p_1;
Hyperparameters are parameters of prior parameters ;
When \alpha = \beta = 1, the distribution is uniform distribution;
To make sure P(p_1 = \frac{1}{6}) is the largest, we can:
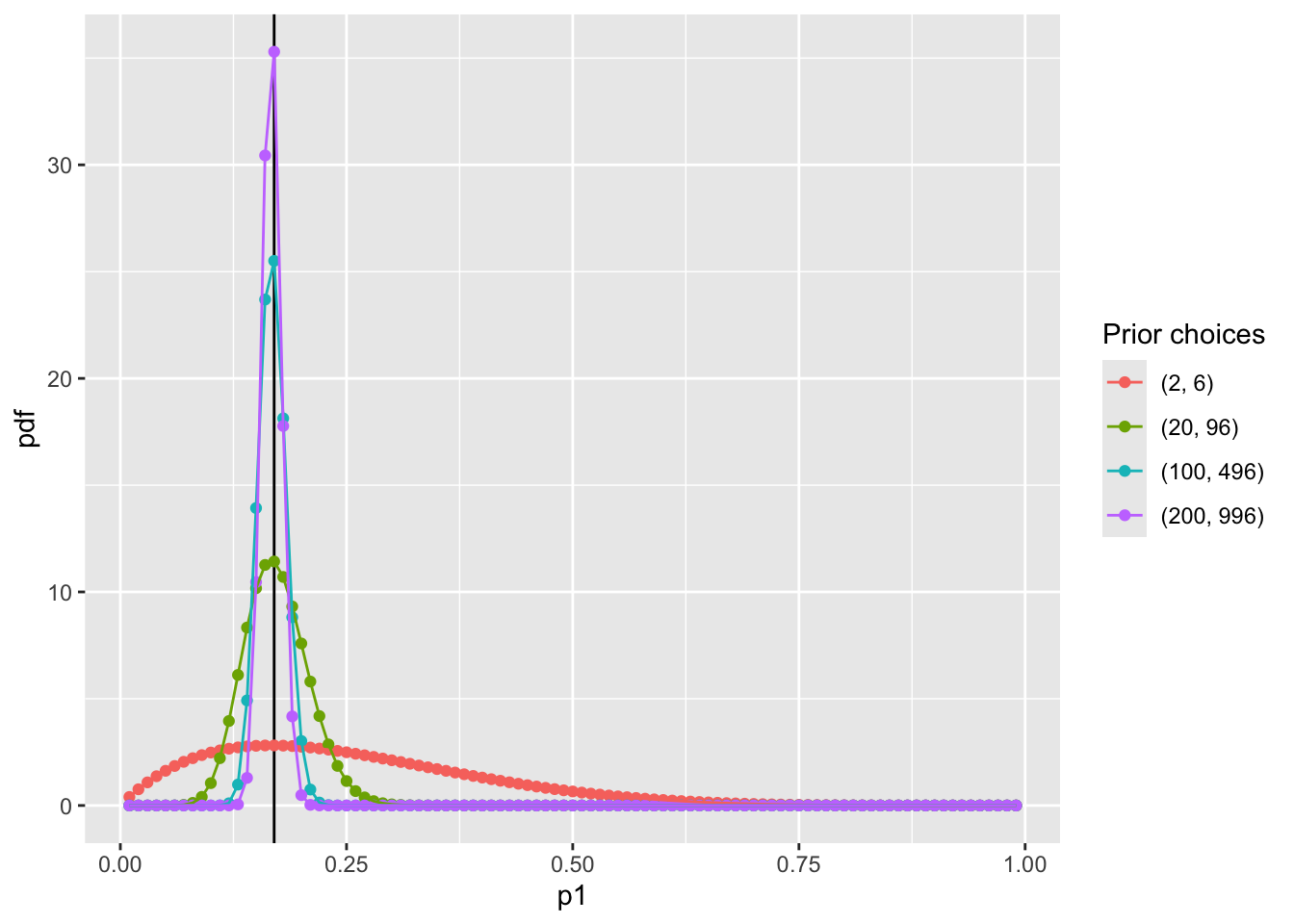
Many choices of prior distribution:
\alpha =2 and \beta = 6 has the same mode as \alpha = 100 and \beta = 496;
hint: when the relationship between \alpha and \eta follows \beta = 5\alpha - 4
The differences between choices is how strongly we feel in our beliefs
8.4 Visualize prior distribution P(p_1)
Click here to see R code
dbeta <- function(p1, alpha, beta) {
# probability density function
PDF = ((p1)^(alpha-1)*(1-p1)^(beta-1)) / beta(alpha, beta)
return(PDF)
}
condition <- data.frame(
alpha = c(2, 20, 100, 200),
beta = c(6, 96, 496, 996)
)
pdf_bycond <- condition %>%
nest_by(alpha, beta) %>%
mutate(data = list(
dbeta(p1 = (1:99)/100, alpha = alpha, beta = beta)
))
## prepare data for plotting pdf by conditions
pdf_forplot <- Reduce(cbind, pdf_bycond$data) %>%
t() %>%
as.data.frame() ## merge conditions together
colnames(pdf_forplot) <- (1:99)/100 # add p1 values as x-axis
pdf_forplot <- pdf_forplot %>%
mutate(Alpha_Beta = c( # add alpha_beta conditions as colors
"(2, 6)",
"(20, 96)",
"(100, 496)",
"(200, 996)"
)) %>%
pivot_longer(-Alpha_Beta, names_to = 'p1', values_to = 'pdf') %>%
mutate(p1 = as.numeric(p1),
Alpha_Beta = factor(Alpha_Beta,levels = unique(Alpha_Beta)))
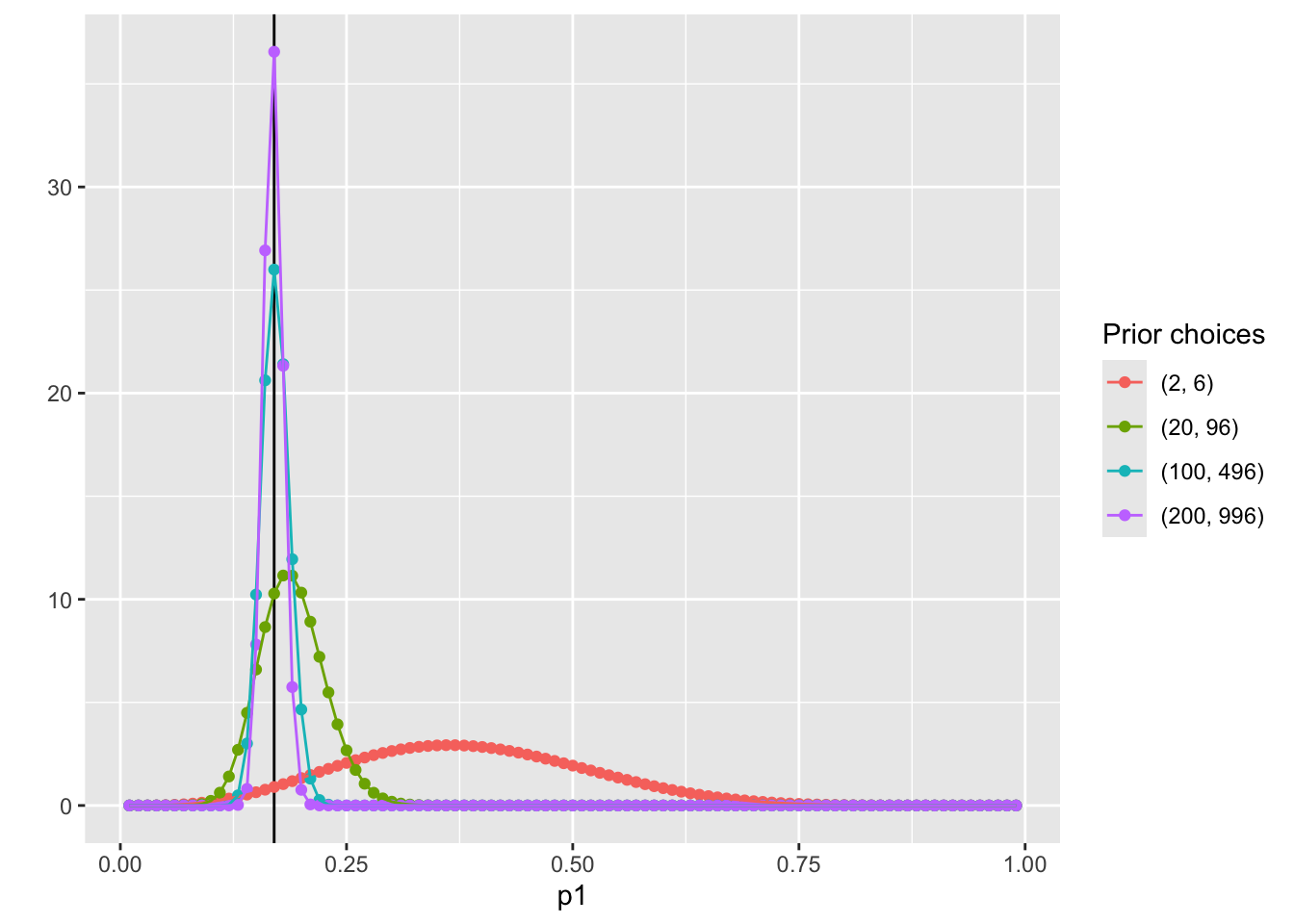
pdf_forplot %>%
ggplot() +
geom_vline(xintercept = 0.17, col = "black") +
geom_point(aes(x = p1, y = pdf, col = Alpha_Beta)) +
geom_path(aes(x = p1, y = pdf, col = Alpha_Beta, group = Alpha_Beta)) +
scale_color_discrete(name = "Prior choices")Question: WHY NOT USE NORMAL DISTRIBUTION?
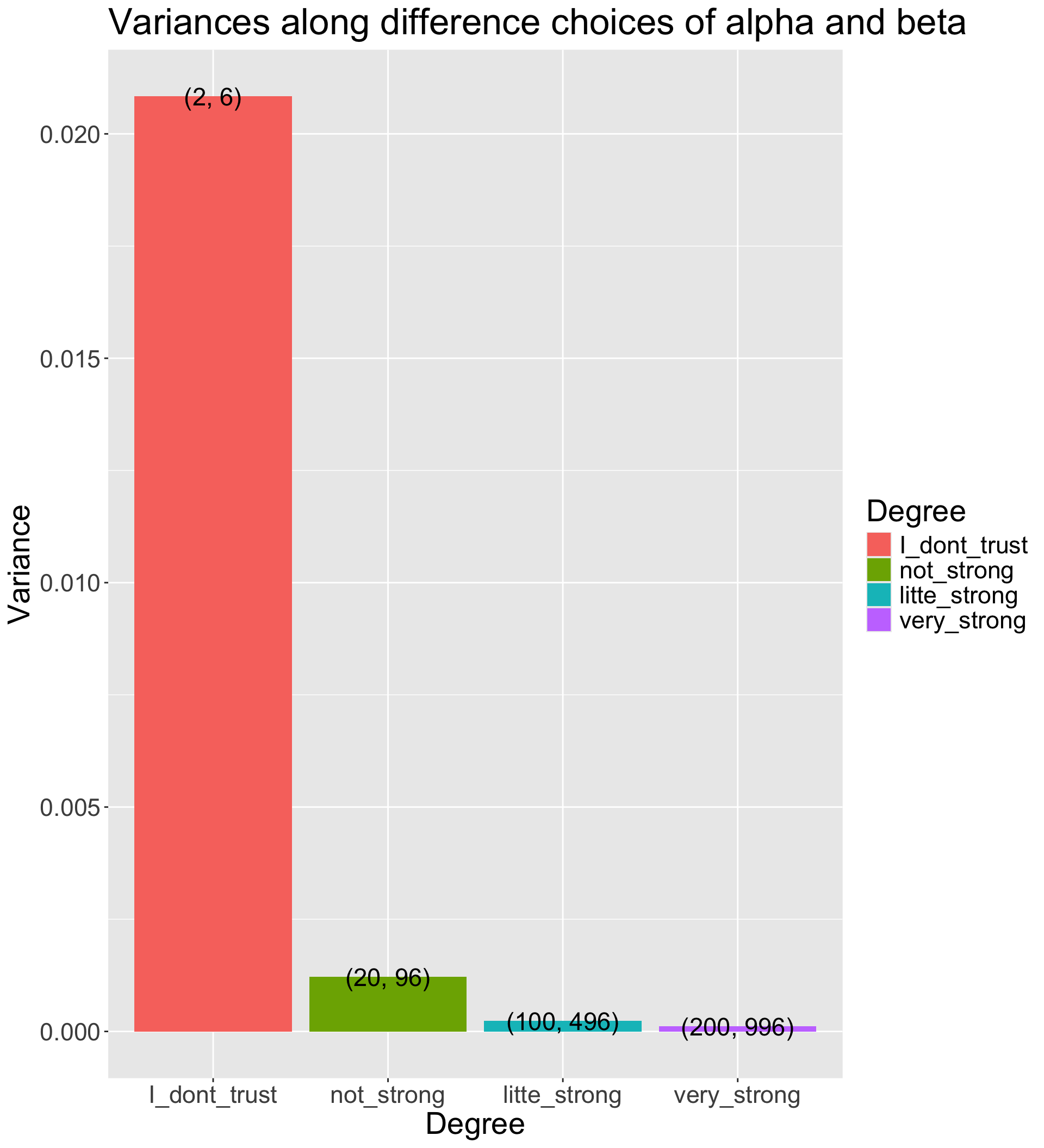
8.5 How strongly we believe in the prior
- The Beta distribution has a variance of \frac{\alpha\beta}{(\alpha+\beta)^2 (\alpha + \beta + 1)}
- The smaller prior variance means the prior is more informative
- Informative priors are those that have relatively small variances
- Uninformative priors are those that have relatively large variances
- Suppose we have four sets of hyperparameters choices: (2,6);(20,96);(100,496);(200,996)
Click here to see R code
# Function for variance of Beta distribution
varianceFun = function(alpha_beta){
alpha = alpha_beta[1]
beta = alpha_beta[2]
return((alpha * beta)/((alpha + beta)^2*(alpha + beta + 1)))
}
# Multiple prior choices from uninformative to informative
alpha_beta_choices = list(
I_dont_trust = c(2, 6),
not_strong = c(20, 96),
litte_strong = c(100, 496),
very_strong = c(200, 996))
## Transform to data.frame for plot
alpha_beta_variance_plot <- t(sapply(alpha_beta_choices, varianceFun)) %>%
as.data.frame() %>%
pivot_longer(everything(), names_to = "Degree", values_to = "Variance") %>%
mutate(Degree = factor(Degree, levels = unique(Degree))) %>%
mutate(Alpha_Beta = c(
"(2, 6)",
"(20, 96)",
"(100, 496)",
"(200, 996)"
))
alpha_beta_variance_plot %>%
ggplot(aes(x = Degree, y = Variance)) +
geom_col(aes(fill = Degree)) +
geom_text(aes(label = Alpha_Beta), size = 6) +
labs(title = "Variances along difference choices of alpha and beta") +
theme(text = element_text(size = 21)) 9 Step 3: The Posterior Distribution
Choose a Beta distribution as the prior distribution of p_1 is very convenient:
When combined with Bernoulli (Binomial) data likelihood, the posterior distribution (P(p_1|Data)) can be derived analytically
The posterior distribution is also a Beta distribution
\alpha' = \alpha + \sum_{i=1}^{N}X_i (\alpha' is parameter of the posterior distribution)
\beta' = \beta + (N - \sum_{i=1}^{N}X_i) (\beta' is parameter of the posterior distribution)
The Beta distribution is said to be a conjugate prior in Bayesian analysis: A prior distribution that leads to posterior distribution of the same family
- Prior and Posterior distribution are all Beta distribution
9.1 Visualize the posterior distribution P(p_1|Data)
Click here to see R code
dbeta_posterior <- function(p1, alpha, beta, data) {
alpha_new = alpha + sum(data)
beta_new = beta + (length(data) - sum(data) )
# probability density function
PDF = ((p1)^(alpha_new-1)*(1-p1)^(beta_new-1)) / beta(alpha_new, beta_new)
return(PDF)
}
# Observed data
dat = c(0, 1, 0, 1, 1)
condition <- data.frame(
alpha = c(2, 20, 100, 200),
beta = c(6, 96, 496, 996)
)
pdf_posterior_bycond <- condition %>%
nest_by(alpha, beta) %>%
mutate(data = list(
dbeta_posterior(p1 = (1:99)/100, alpha = alpha, beta = beta,
data = dat)
))
## prepare data for plotting pdf by conditions
pdf_posterior_forplot <- Reduce(cbind, pdf_posterior_bycond$data) %>%
t() %>%
as.data.frame() ## merge conditions together
colnames(pdf_posterior_forplot) <- (1:99)/100 # add p1 values as x-axis
pdf_posterior_forplot <- pdf_posterior_forplot %>%
mutate(Alpha_Beta = c( # add alpha_beta conditions as colors
"(2, 6)",
"(20, 96)",
"(100, 496)",
"(200, 996)"
)) %>%
pivot_longer(-Alpha_Beta, names_to = 'p1', values_to = 'pdf') %>%
mutate(p1 = as.numeric(p1),
Alpha_Beta = factor(Alpha_Beta,levels = unique(Alpha_Beta)))
pdf_posterior_forplot %>%
ggplot() +
geom_vline(xintercept = 0.17, col = "black") +
geom_point(aes(x = p1, y = pdf, col = Alpha_Beta)) +
geom_path(aes(x = p1, y = pdf, col = Alpha_Beta, group = Alpha_Beta)) +
scale_color_discrete(name = "Prior choices") +
labs( y = '')9.2 Manually calculate summaries of the posterior distribution
To determine the estimate of p_1, we need to summarize the posterior distribution:
With prior hyperparameters \alpha = 2 and \beta = 6:
\hat p_1 = \frac{2+3}{2+3+6+2}=\frac{5}{13} = .3846
SD = 0.1300237
With prior hyperparameters \alpha = 100 and \beta = 496:
\hat p_1 = \frac{100+3}{100+3+496+2}=\frac{103}{601} = .1714
SD = 0.0153589
uninfo <- pdf_posterior_forplot %>% filter(Alpha_Beta == "(2, 6)")
paste0("Sample Posterior Mean: ", mean(uninfo$p1 * uninfo$pdf)) # .3885[1] "Sample Posterior Mean: 0.388500388484552"uninfo2 <- pdf_posterior_forplot %>% filter(Alpha_Beta == "(100, 496)")
paste0("Sample Posterior Mean: ", mean(uninfo2$p1 * uninfo2$pdf)) # 0.1731[1] "Sample Posterior Mean: 0.173112153145432"9.3 Stan Code for the example model
library(cmdstanr)
set_cmdstan_path("~/cmdstan/")
register_knitr_engine(override = FALSE)data {
int<lower=0> N; // sample size
array[N] int<lower=0, upper=1> y; // observed data
real<lower=1> alpha; // hyperparameter alpha
real<lower=1> beta; // hyperparameter beta
}
parameters {
real<lower=0,upper=1> p1; // parameters
}
model {
p1 ~ beta(alpha, beta); // prior distribution
for(n in 1:N){
y[n] ~ bernoulli(p1); // model
}
}Click here to see R code
# mod <- cmdstan_model('dice_model.stan')
data_list1 <- list(
y = c(0, 1, 0, 1, 1),
N = 5,
alpha = 2,
beta = 6
)
data_list2 <- list(
y = c(0, 1, 0, 1, 1),
N = 5,
alpha = 100,
beta = 496
)
fit1 <- dice_model$sample(
data = data_list1,
seed = 123,
chains = 4,
parallel_chains = 4,
refresh = 1000
)Running MCMC with 4 parallel chains...
Chain 1 Iteration: 1 / 2000 [ 0%] (Warmup)
Chain 1 Iteration: 1000 / 2000 [ 50%] (Warmup)
Chain 1 Iteration: 1001 / 2000 [ 50%] (Sampling)
Chain 1 Iteration: 2000 / 2000 [100%] (Sampling)
Chain 2 Iteration: 1 / 2000 [ 0%] (Warmup)
Chain 2 Iteration: 1000 / 2000 [ 50%] (Warmup)
Chain 2 Iteration: 1001 / 2000 [ 50%] (Sampling)
Chain 2 Iteration: 2000 / 2000 [100%] (Sampling)
Chain 3 Iteration: 1 / 2000 [ 0%] (Warmup)
Chain 3 Iteration: 1000 / 2000 [ 50%] (Warmup)
Chain 3 Iteration: 1001 / 2000 [ 50%] (Sampling)
Chain 3 Iteration: 2000 / 2000 [100%] (Sampling)
Chain 4 Iteration: 1 / 2000 [ 0%] (Warmup)
Chain 4 Iteration: 1000 / 2000 [ 50%] (Warmup)
Chain 4 Iteration: 1001 / 2000 [ 50%] (Sampling)
Chain 4 Iteration: 2000 / 2000 [100%] (Sampling)
Chain 1 finished in 0.0 seconds.
Chain 2 finished in 0.0 seconds.
Chain 3 finished in 0.0 seconds.
Chain 4 finished in 0.0 seconds.
All 4 chains finished successfully.
Mean chain execution time: 0.0 seconds.
Total execution time: 0.2 seconds.Click here to see R code
fit2 <- dice_model$sample(
data = data_list2,
seed = 123,
chains = 4,
parallel_chains = 4,
refresh = 1000
)Running MCMC with 4 parallel chains...
Chain 1 Iteration: 1 / 2000 [ 0%] (Warmup)
Chain 1 Iteration: 1000 / 2000 [ 50%] (Warmup)
Chain 1 Iteration: 1001 / 2000 [ 50%] (Sampling)
Chain 1 Iteration: 2000 / 2000 [100%] (Sampling)
Chain 2 Iteration: 1 / 2000 [ 0%] (Warmup)
Chain 2 Iteration: 1000 / 2000 [ 50%] (Warmup)
Chain 2 Iteration: 1001 / 2000 [ 50%] (Sampling)
Chain 2 Iteration: 2000 / 2000 [100%] (Sampling)
Chain 3 Iteration: 1 / 2000 [ 0%] (Warmup)
Chain 3 Iteration: 1000 / 2000 [ 50%] (Warmup)
Chain 3 Iteration: 1001 / 2000 [ 50%] (Sampling)
Chain 3 Iteration: 2000 / 2000 [100%] (Sampling)
Chain 4 Iteration: 1 / 2000 [ 0%] (Warmup)
Chain 4 Iteration: 1000 / 2000 [ 50%] (Warmup)
Chain 4 Iteration: 1001 / 2000 [ 50%] (Sampling)
Chain 4 Iteration: 2000 / 2000 [100%] (Sampling)
Chain 1 finished in 0.0 seconds.
Chain 2 finished in 0.0 seconds.
Chain 3 finished in 0.0 seconds.
Chain 4 finished in 0.0 seconds.
All 4 chains finished successfully.
Mean chain execution time: 0.0 seconds.
Total execution time: 0.2 seconds.9.3.1 Uninformative prior distribution (2, 6) and posterior distribution
Click here to see R code
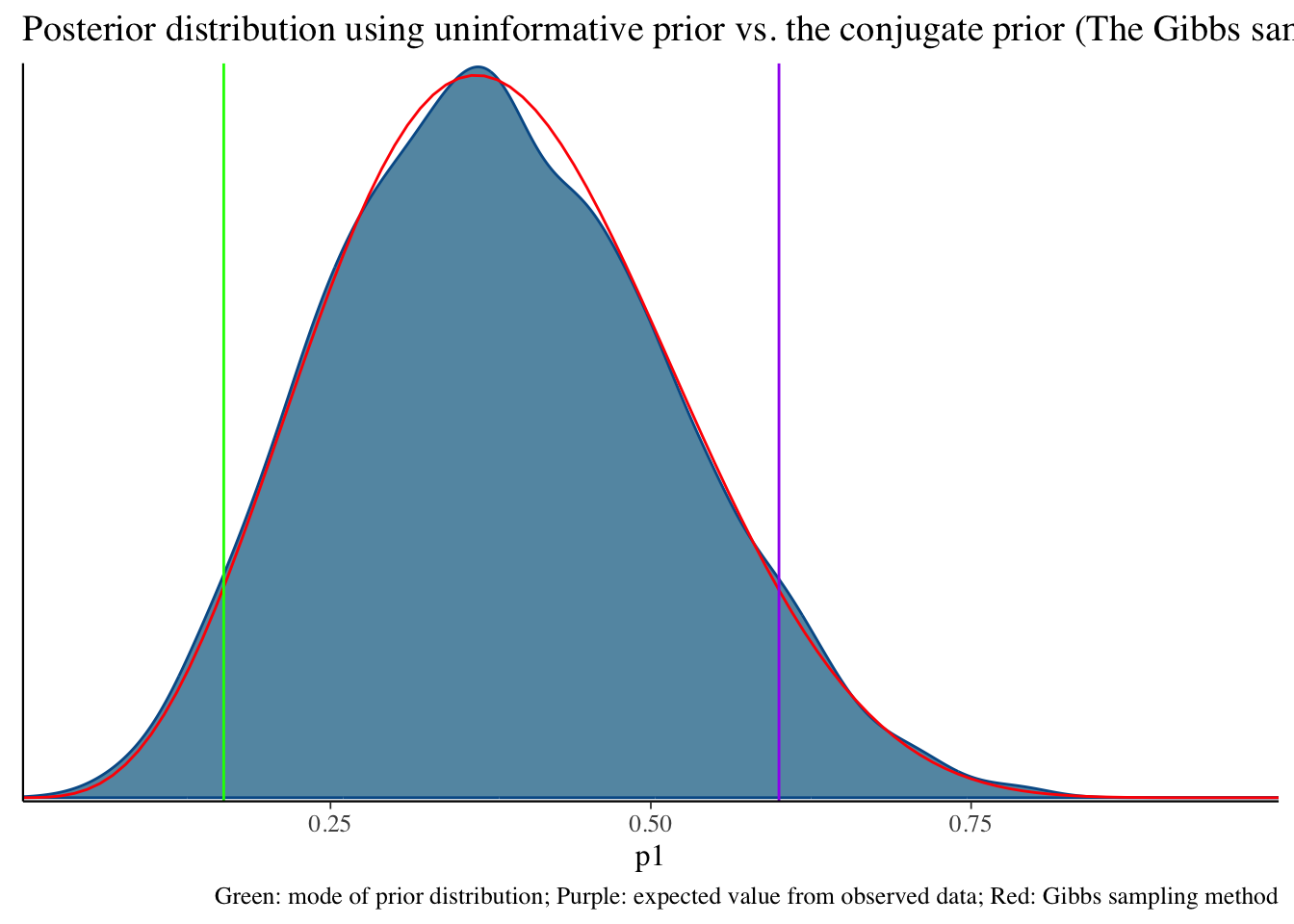
bayesplot::mcmc_dens(fit1$draws('p1')) +
geom_path(aes(x = p1, y = pdf), data = pdf_posterior_forplot %>% filter(Alpha_Beta == '(2, 6)'), col = "red") +
geom_vline(xintercept = 1/6, col = "green") +
geom_vline(xintercept = 3/5, col = "purple") +
labs(title = "Posterior distribution using uninformative prior vs. the conjugate prior (The Gibbs sampler) method", caption = 'Green: mode of prior distribution; Purple: expected value from observed data; Red: Gibbs sampling method')Click here to see R code
fit1$summary()# A tibble: 2 × 10
variable mean median sd mad q5 q95 rhat ess_bulk ess_tail
<chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 lp__ -9.19 -8.91 0.751 0.337 -10.7 -8.66 1.00 1812. 2400.
2 p1 0.383 0.377 0.131 0.137 0.180 0.609 1.00 1593. 1629.9.3.2 Informative prior distribution (100, 496) and posterior distribution
Click here to see R code
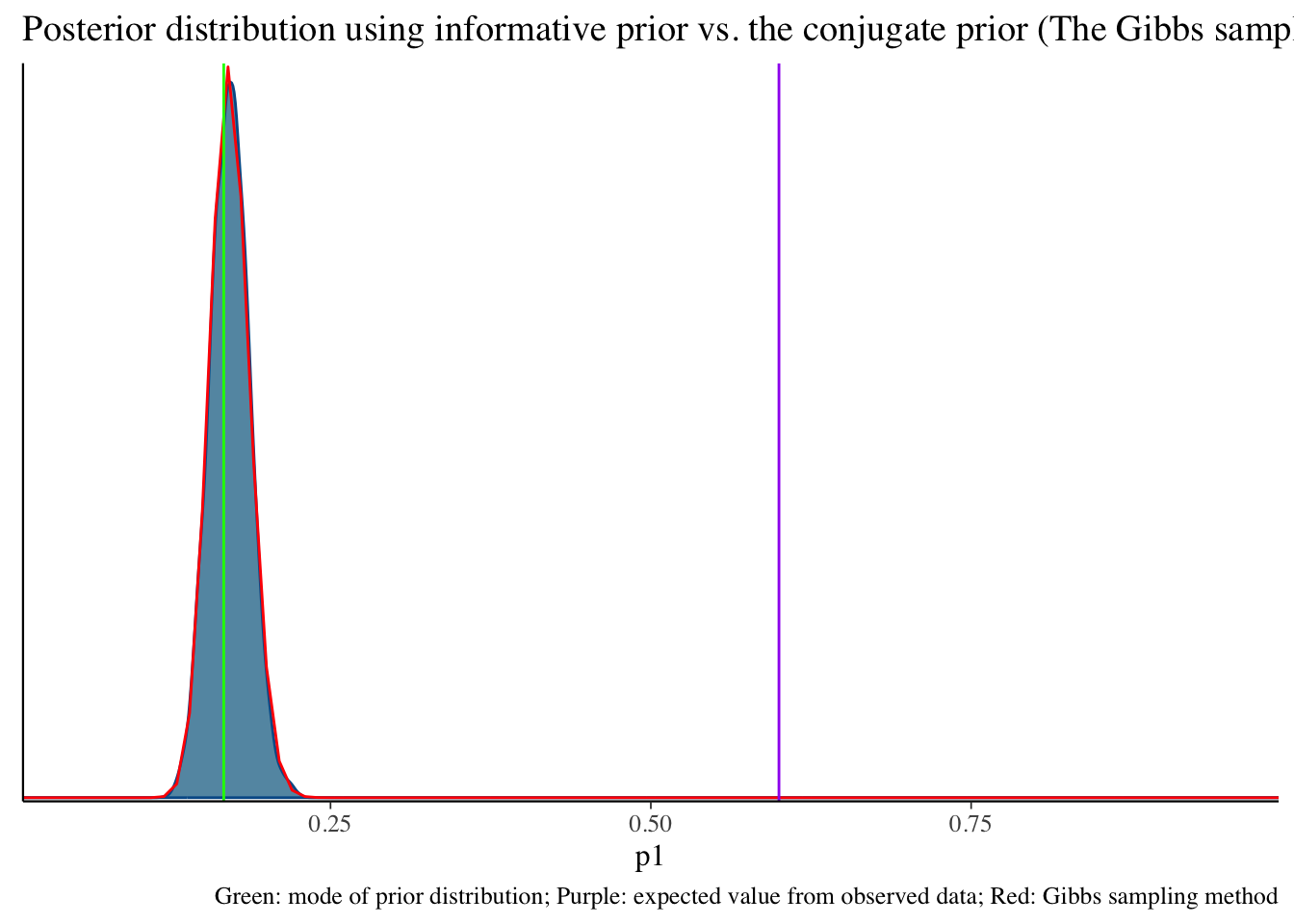
bayesplot::mcmc_dens(fit2$draws('p1')) +
geom_path(aes(x = p1, y = pdf), data = pdf_posterior_forplot %>% filter(Alpha_Beta == '(100, 496)'), col = "red") +
geom_vline(xintercept = 1/6, col = "green") +
geom_vline(xintercept = 3/5, col = "purple") +
labs(title = "Posterior distribution using informative prior vs. the conjugate prior (The Gibbs sampler) method", caption = 'Green: mode of prior distribution; Purple: expected value from observed data; Red: Gibbs sampling method')Click here to see R code
fit2$summary()# A tibble: 2 × 10
variable mean median sd mad q5 q95 rhat ess_bulk
<chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 lp__ -276. -276. 0.716 0.317 -277. -275. 1.00 1816.
2 p1 0.171 0.171 0.0154 0.0155 0.146 0.197 1.00 1627.
# ℹ 1 more variable: ess_tail <dbl>10 Exercise
Install cmdstan and cmdstanr
Practice beta prior distribution with hyperparameter {2, 3}, {10, 15}
Summarize the results
11 Next Class
We will talk more about how Bayesian model works. Thank you.
12 Bonus: Conjugacy of Beta Distribution
When the binomial likelihood is multiplied by the beta prior, the result is proportional to another beta distribution:
- \text{Posterior} \propto \theta^x (1 - \theta)^{n-x} \times \theta^{\alpha - 1} (1 - \theta)^{\beta - 1}
- Simplifies to: \theta^{x + \alpha - 1} (1 - \theta)^{n - x + \beta - 1}
This is the kernel of a beta distribution with updated parameters \alpha' = x + \alpha and \beta' = n - x + \beta. The fact that the posterior is still a beta distribution is what makes the beta distribution a conjugate prior for the binomial likelihood.
Human langauge: Both beta (prior) and binomial (likelihood) are so-called “exponetial family”. The muliplication of them is still a “exponential family” distribution.
13 Bayesian updating
We can use the posterior distribution as a prior!
1. data {0, 1, 0, 1, 1} with the prior hyperparameter {2, 6} -> posterior parameter {5, 9}
2. new data {1, 1, 1, 1, 1} with the prior hyperparameter {5, 9} -> posterior parameter {10, 9}
3. one more new data {0, 0, 0, 0, 0} with the prior hyperparameter {10, 9} -> posterior parameter {10, 14}
14 Wrapping up
Today is a quick introduction to Bayesian Concept
- Bayes’ Theorem: Fundamental theorem of Bayesian Inference
- Prior distribution: What we know about the parameter before seeing the data
- hyperparameter: parameter of the prior distribution
- Uninformative prior: Prior distribution that does not convey any information about the parameter
- Informative prior: Prior distribution that conveys information about the parameter
- Conjugate prior: Prior distribution that makes the posterior distribution the same family as the prior distribution
- Likelihood: What the data tell us about the parameter
- Likelihood function: Probability of the data given the parameter
- Likelihood principle: All the information about the data is contained in the likelihood function
- Likelihood is a sort of “scientific judgment” on the generation process of the data
- Posterior distribution: What we know about the parameter after seeing the data